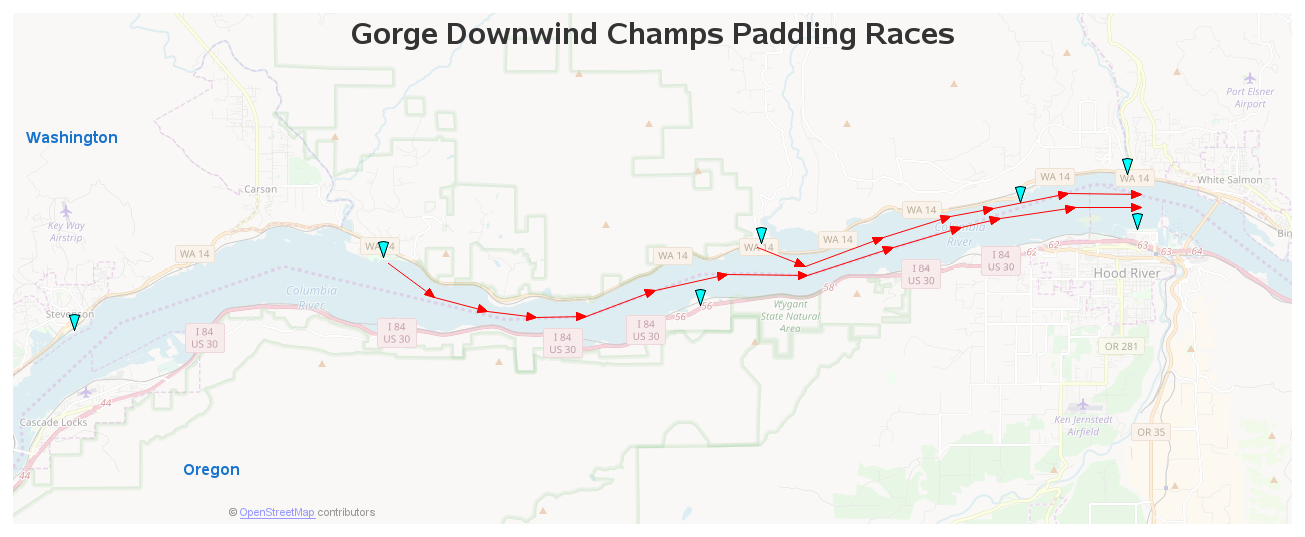
  Or with the same twoway command, I might get any of the following graphs. The one exception is the transparency in the scatterplot markers and confidence interval area I requested using %8 and %20 within the color() option. twoway scatter observed1 observed2 day, color(%8 %8) || To demonstrate, I use a graph with overlaid scatterplots, model fit lines, and a confidence interval. In any case, you start with a graph of your data or results, and you need to transform that graph into the style you want. You want a graph with colors that everyone can differentiate. You want a graph that fits the style of your journal. Geom_sf(data = pts_random, color = "#F8766D",size=0.You want a graph that most effectively communicates your message. # (alternatively replace pts_random by a real sampling dataset (see Meyer and Pebesma 2022): # (refer to figure in Meyer and Pebesma 2022) Type = "feature",variables=c("DEM","TWI", "NDRE.M"))ĭist <- plot_geodist(x=pts_train, modeldomain=studyArea, cvfolds = folds, testdata = pts_test, Plot_geodist(x=pts_train, modeldomain=studyArea, # Distance between training data, new data and CV folds:įolds <- createFolds(1:nrow(pts_train),k=3,returnTrain=FALSE)ĭist <- plot_geodist(x=pts_train, modeldomain=studyArea, cvfolds=folds)ĭist <- plot_geodist(x=pts_train, modeldomain=studyArea,Ĭvfolds=nndm_pred$indx_test, cvtrain=nndm_pred$indx_train) #mapview(pts_train,col.regions="blue")+mapview(pts_test,col.regions="red")ĭist <- plot_geodist(pts_train,studyArea,testdata=pts_test) # Distance between training data, new data and test data: # Distance between training data and new data:ĭist <- plot_geodist(pts_train,studyArea) StudyArea <- raster::stack(system.file("extdata","predictors_.grd",package="CAST")) Pts <- st_as_sf(dat,coords=c("Easting","Northing")) Hanna Meyer, Edzer Pebesma, Marvin Ludwigĭat <- get(load(system.file("extdata","Cookfarm.RData",package="CAST")))ĭat <- aggregate(dat,īy=list(as.character(dat$SOURCEID)),mean) See Meyer and Pebesma (2022) for an application of this plotting function Unit of returned geographic distances is meters. If not provided they are extracted from the modeldomain rasterStack.Ī list including the plot and the corresponding ame containing the distances. Predictor values for x are optional if modeldomain is a raster. If type = "feature", the argument modeldomain (and if provided then also the testdata) has to include predictors. 
The function takes a regular point sample (amount defined by samplesize) from the spatial extent. The modeldomain is a sf polygon or a raster that defines the prediction area. "density" for density plot or "ecdf" for cumulative plot. Only if type="geo" and only applied to the plot. If not provided all variables included in modeldomain are used.Ĭharacter. Use sampling = "Fibonacci" for global applications.Ĭharacter vector defining the predictor variables used if type="feature. #Compute geodist for each row stata how toHow to draw prediction samples? See spsample. How many prediction samples should be used?Ĭharacter. object of class sf: Data used for independent validation If cvtrain is null but cvfolds is not, all samples but those included in cvfolds are used as training data List of row indices of x to fit the model to in each CV iteration. List of row indices of x that are held back in each CV iteration. Should the distance be computed in geographic space or in the normalized multivariate predictor space (see Details) Raster or sf object defining the prediction area (see Details) 
Object of class sf, training data locations Alternatively distances can also be calculated in the multivariate feature space. The plot can be used to check the suitability of a chosen CV method to be representative to estimate map accuracy. Optional, the nearest neighbor distances between training data and test data or between training data and CV iterations is shown. Plot euclidean nearest neighbor distances in geographic space or feature space Descriptionĭensity plot of nearest neighbor distances in geographic space or feature space between training data as well as between training data and prediction locations. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
